Improving Patient Outcomes with Stewardship and Rapid Molecular Diagnostics
A recent analysis of several studies found that when molecular rapid diagnostic tests (RDTs) are combined with antimicrobial stewardship (AMS) programs, patient outcomes are improved exponentially. When combined with AMS interventions, molecular RDTs decreased mortality by 36% in patients with bloodstream infections and reduced both the time to effective therapy by 5.03 hours and hospital length of stay by 2.48 days.
In addition to improving patient outcomes, RDTs may help AMS programs achieve aspects of the CDC’s Core Elements of Antimicrobial Stewardship by:
- Ensuring prospective audit and feedback of certain RDT results
- Educating staff through AMS interpretation and intervention based on RDT results
- Tracking/monitoring by assessment of mortality and hospital length of stay outcomes
The analysis concludes that RDTs link AMS programs and clinical microbiology labs in a mutually beneficial relationship and notes the important role pharmacists will play in helping labs translate the undeniable clinical value of RDTs into financial cost-justifications.
Advances in RDTs provide new opportunities for AMS and enhance the function of clinical microbiology laboratories, but diagnostic stewardship will be crucial going forward as the clinical benefit of RDTs has been solidified. Pharmacists must make a concerted effort to justify the addition of RDTs prior to their implementation, educate prescribing staff of the benefits, and track RDT-related outcomes within the institution."
Assistant Professor, University of Illinois at Chicago
Wenzler E. Using rapid diagnostics to fight resistance. Infectious Disease Special Edition. May 30, 2018. https://www.idse.net/Resistance--Stewardship/Article/09-15/Using-Rapid-Diagnostics-to-Fight-Resistance/48730
CDC Highlights Need for Active CRE Surveillance to Combat Antibiotic Resistance
According to the Centers for Disease Control and Prevention (CDC), “nightmare bacteria” that have the power to resist most or all antibiotics, are popping up across the U.S. However, new, aggressive policies, which have been shown to be effective, can help stop them from spreading.1 The CDC’s containment policy includes active surveillance with rapid detection tests to reduce the spread of antibiotic resistance.2
The CDC’s new program for testing suspect bacteria turned up unusual antibiotic-resistant genes 221 times in 2017, and 11% of people screened for these superbugs carried them even though they had no symptoms. The CDC tested for two of the best-known superbug germs: carbapenem-resistant Enterobacteriaceae (CRE) and carbapenem-resistant Pseudomonas aeruginosa bacteria.1
Antibiotic-resistant germs kill more than 23,000 Americans a year, and the World Health Organization has labeled antibiotic resistance a “fundamental threat” to humanity.3 Dr. Anne Schuchat, Acting Principal Deputy Director of CDC, reported that the CDC estimated that even if the containment policy were only 20% effective, it could reduce the number of nightmare bacteria cases by 76% over three years in one area.2 The policy requires a coordinated response among health care facilities, labs, and health departments.1
What the CDC is highlighting is the need, once you identify people carrying these bugs, to screen them and also their contacts to make sure they haven’t passed these bugs."
Director of the Antibiotic Resistance Action Center at George Washington University
References:
1. https://www.nbcnews.com/health/health-news/nightmare-bacteria-are-trying-spread-u-s-cdc-says-n862436
2. Containment of Novel Multidrug-Resistant Organisms and Resistance Mechanisms — United States, 2006–2017. https://www.cdc.gov/mmwr/volumes/67/wr/mm6713e1.htm?s_cid=mm6713e1_e
3. https://www.nbcnews.com/health/health-news/who-labels-drug-resistant-superbugs-fundamental-threat-humans-n651981
Partner with Your Laboratory — ASP Improvements with Rapid Diagnostics
Precise diagnosis is one of the cornerstones of a successful antimicrobial stewardship program (ASP), as reviewed by experts from the University of Montreal and the University of Chicago in a recent publication.1 Partnering with your laboratory to implement high-performing rapid diagnostic tests such as nucleic acid amplification tests can reduce the inappropriate use of antimicrobials and contribute to improvements in your stewardship program.
Click here to download the study to explore a variety of stewardship concepts and how successful partnerships between the lab and clinicians can play a vital role in your stewardship program.
1. Morency-Potvin P, Schwartz DN, Weinstein RA. 2017. Antimicrobial stewardship: how the microbiology laboratory can right the ship. Clin Microbiol Rev 30:381– 407. https://doi.org/10.1128/CMR.00066-16. Downloaded February 1, 2018 from http://cmr.asm.org/
Limiting Inappropriate Antibiotic Use in the Emergency Department with the Help of PCR
Watch Now
Dr. Larissa May discusses how clinical practice guidelines on skin and soft tissue infections make it more important than ever to use diagnostics to guide targeted antibiotic use. Hear how her institution was able to reduce inappropriate antibiotic use in the ED with the help of on-demand PCR testing for MRSA and S. aureus.
For in vitro Diagnostic UseTEDx Talk Brings Antibiotic Stewardship Message to the Masses
Watch Now
In a recent TEDx talk, Dr. Debbie Goff, a clinical pharmacist and global antimicrobial stewardship advocate, delivers a compelling case for shared responsibility in the fight against drug-resistant organisms. With real-life examples of “just in case” antibiotic use, the case for stewardship is presented in a relatable format suitable for clinicians and non-clinicians alike.
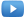
Chlorhexidine Use Can Lead to Colistin Resistance
A recently published scientific study identifies a novel resistance mechanism to the widely used antiseptic, chlorhexidine, which fosters cross-resistance to Colistin, an antibiotic of last resort for treatment of carbapenem-resistant infections. The study shows that mutations conferring chlorhexidine resistance in Klebsiella pneumoniae lead to generation of mutations that result in Colistin resistance (Figure 1). Interestingly, this appears to be unidirectional − Colistin resistance does not lead to chlorhexidine resistance.
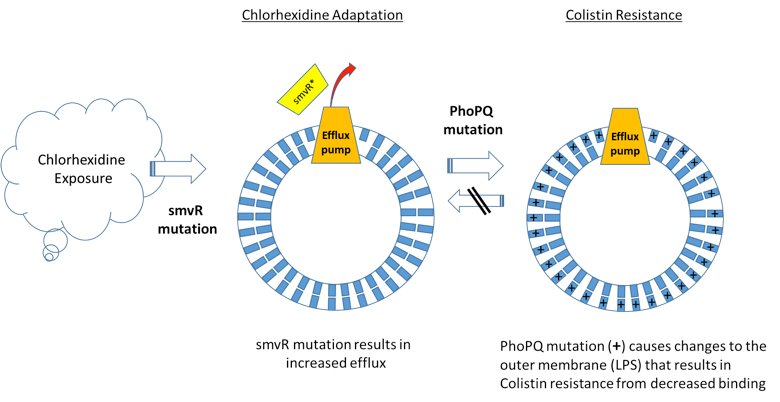
Based on these findings, eradicating Colistin-resistant Klebsiella isolates may become increasingly problematic given the current and increasingly broad use of chlorhexidine. While the study affirms Darwin − evolutionary pressure leads to the development of new resistance mechanisms – loss of an antibiotic of last resort like Colistin has wide-ranging and potentially lethal implications, including prolonging of outbreaks or leading to new outbreaks. This article highlights the challenges and should generate discussion about common infection control practices using chlorhexidine to prevent healthcare associated infections. Employing more vigilant anti-microbial resistance (AMR) monitoring, including sensitive and specific molecular diagnostics as early surveillance, may be one strategy to address the dilemma posed by these results and reduce the potential for AMR infections.
Click here for more on the study from Antimicrobial Agents and Chemotherapy.
Wand ME, Bock JL, Bonney LC, Sutton JM. Mechanisms of increased resistance to chlorhexidine and cross-resistance to colistin following exposure of Klebsiella pneumoniae clinical isolates to chlorhexidine. Antimicrob Agents Chemother. 2016 Oct 31. doi: 10.1128/AAC.01162-16 [Epub ahead of print].
Reduced empiric antibiotic therapy with rapid molecular testing for C. difficile
Compared with traditional testing methods*, on-demand PCR testing with Xpert C. difficile resulted in:
- 20% fewer patients receiving empiric antibiotic therapy
- 62% reduction in days of empiric therapy (2.3 vs 0.9 days; P = 0.007)
- 3.33 fewer doses per patient administered
- 42% fewer tests performed (per patient)
- No increase in total cost of care (despite higher price-per-test)
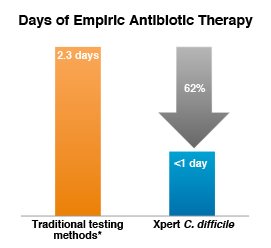
* C. DIFFICILE TOX A/B II EIA Test (TechLab).
Peppard WJ and Ledeboer NA. Implementation of Polymerase Chain Reaction to Rule Out Clostridium difficile Infection Is Associated With Reduced Empiric Antibiotic Duration of Therapy. Hosp Pharm. 2014;49:639-43.
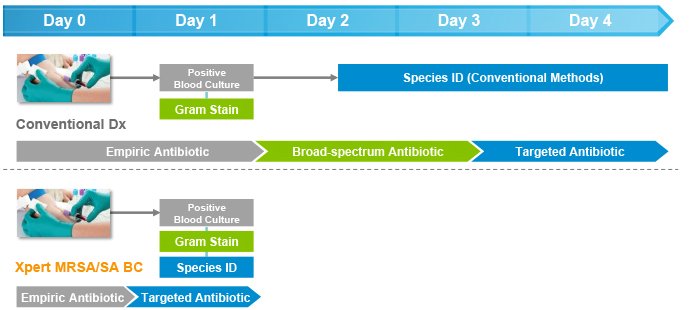
Faster detection of MRSA and S. aureus from positive blood cultures optimizes treatment decisions & outcomes
On-demand molecular diagnostic testing with Cepheid's Xpert MRSA/SA BC plus antimicrobial stewardship intervention resulted in a reduction in:
- Mean time to optimal antibiotic therapy of 1.7 days
- Mean length of stay of 6.2 days
- Mean hospital costs of $21,387 per patient
Bauer KA, West JE, Balada-Llasat JM, et al. An antimicrobial stewardship program's impact with rapid polymerase chain reaction methicillin-resistant Staphylococcus aureus/S. aureus blood culture test in patients with S. aureus bacteremia. Clin Infect Dis. 2010 1;51:1074-80.








